All the features are displayed on the map using user-modifiable colors the features can be manually entered, imported from GenBank or automatically found after scanning by Serial Cloner. The graphic map of Serial Cloner is really Graphic as you can easily select and extract a fragment or show single, double or multiple cutter all in the same window. Since version 2.1, Sequence Features are visible in the Sequence Map. Serial Cloner also lets you build text restriction map and quickly format it to add multi-frame translation or only show single cutters for example. Numerically select fragments, find restriction sites, ORF or any nucleotide or peptide sequence, calculate Tm of selected fragments, %GC or dynamically determine the translation your selection into peptide and calculate the MW using a compact interface. Serial Cloner 2.5, handles Annotations and Features both in the sequence and in the Graphic Map and can automatically scan for sequence Features.Īll the tools you need to analyze and manipulate your sequences are available in an all-in-one-window concept. Powerful graphical display tools and simple interfaces help the analysis and construction steps in a very intuitive way. 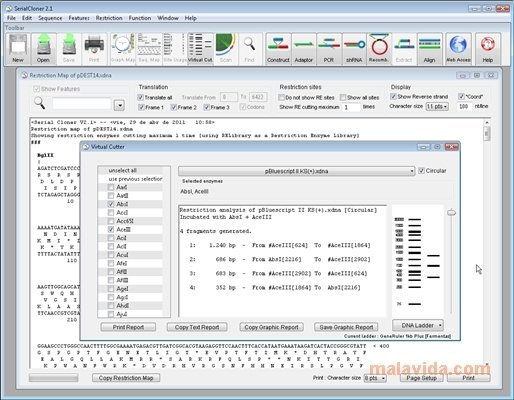
Import from VectorNTI multi-file format is also supported. Serial Cloner also import files saved in the Vector NTI, MacVector, ApE, DNAstar, pDRAW32 and GenBank formats. Serial Cloner reads and write DNA Strider-compatible files and import and export files in the universal FASTA format. Serial Cloner has been developed to provide a light yet powerful molecular biology software to both Macintosh and Windows users. Serial Cloner also allows you to quickly prepare the text file in Fasta format, or send a request for a multi-alignment in Clustal.Serial Cloner has been developed to provide a light yet powerful molecular biology software to its users. Finally, Serial Cloner provides an interface to align two sequences by using an algorithm or a local server and retrieve the boat NCBI BLAST2Seq virtual consensus, a web browser with instant import records from NCBI / EMBL and quiet of the restriction map of the generator. Optional interface allows easy Gateway (TM) clone for BP and LR reactions. Just choose a blunt if you need, and click bandage. You can collect the fragments obtained by PCR synthesis adapter / shRNA or simply graphically selecting fragments between restriction sites. shRNA design based on predetermined scaffolding also automated. In addition to the classical restriction maps - both graphic and text - or windows using this site, you can quickly adjust the gain in a PCR-based or prepare summaries. Serial Cloner will help in the creation of new sub-cloning projects and in the preparation of electronic versions of your designs. Graphics Card Serial Cloner is really Graphic as you can easily select and fragment function extract or show one, two or more cutter all in the same window. Serial Cloner also allows you to create text and format restriction map to add a multi-mode translation or only show single cutters for example in a hurry. All the main functions are available through a compact interface. All the tools needed to analyze and manipulate sequences are available in the concept of an all-in-one-window. Powerful graphical mapping tools and simple interfaces help the analysis and construction in a very intuitive way. 
File GenBank / EMBL can be read or imported from the Internet using the included web interface. Furthermore, since the version it has the ability to read and APE files VectorNTI with their capabilities, as well as sequences pDRAW32. Serial Cloner can read and write DNA Strider-compatible files, and import or export files in FASTA. You can comment on any part of the sequence, or use the automatic scan to detect similarities. Serial Cloner now handles Features / Annotations, which has been the most requested improvement. Serial Cloner is a Molecular Biology software that provides tools with an intuitive interface to help you in DNA cloning, sequence analysis and visualization.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
